
Just over three years ago we launched the first Etch A Cell project (https://www.zooniverse.org/projects/h-spiers/etch-a-cell). The project was the first of its kind on the Zooniverse: never before had we asked volunteers to help draw around the small structures inside of cells (also known as ‘manual segmentation of organelles’) visualised with very high-powered electron microscopes. We even had to develop a new tool type on the Zooniverse to do this – a drawing tool for annotating images.
In this first Etch A Cell project, the organelle we asked Zooniverse volunteers to help examine was the nuclear envelope (as you can see shown in green in the image below). The nuclear envelope is a large membrane found within cells. It surrounds the nucleus, which is the part of the cell that contains the genetic material. It’s an important structure to study as it’s known to be involved in a number of diseases, including cancer, and it’s often the first structure research teams inspect in a new data set.

The results…
Earlier this year, we published the first set of results from this project. I’ve summarised some of our most exciting findings below, but if you’d like to take a look at the original paper, you can access it here (https://www.biorxiv.org/content/10.1101/2020.07.28.223024v1.full).
1. Zooniverse volunteers dedicated a huge amount of effort! Zooniverse volunteers submitted more than 100,000 segmentations across the 4000 images analysed in this first Etch A Cell project. Through this effort, the nuclear envelopes of 18 cells were segmented (shown below in green) from our original data block (shown below).
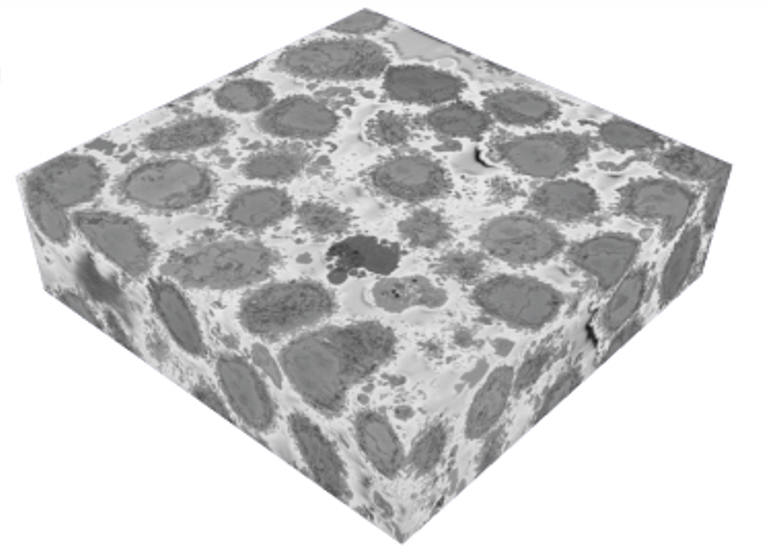
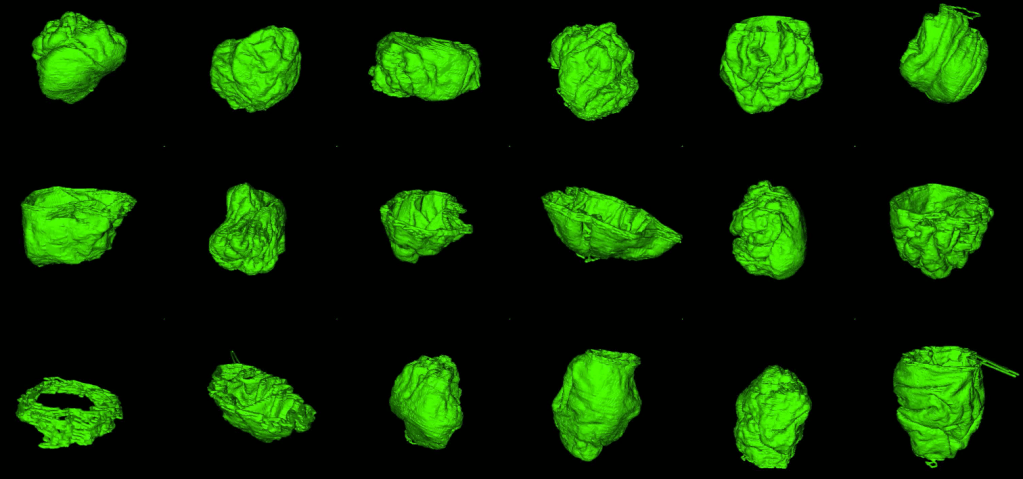

2. Volunteers were very good at segmenting the nuclear envelope. As you can see in the gif and images below, most classifications submitted for each image were really good! Manual segmentation isn’t an easy task to do, even for experts, so we were really impressed!


3. There’s power in a crowd! The image below shows an overlay of every single segmentation for one of the nuclei studied in Etch A Cell. As you can see, through the collective effort of Zooniverse volunteers, something beautiful emerges – by overlaying everyone’s effort like this, you can see the shape of the nuclear envelope begin to appear!
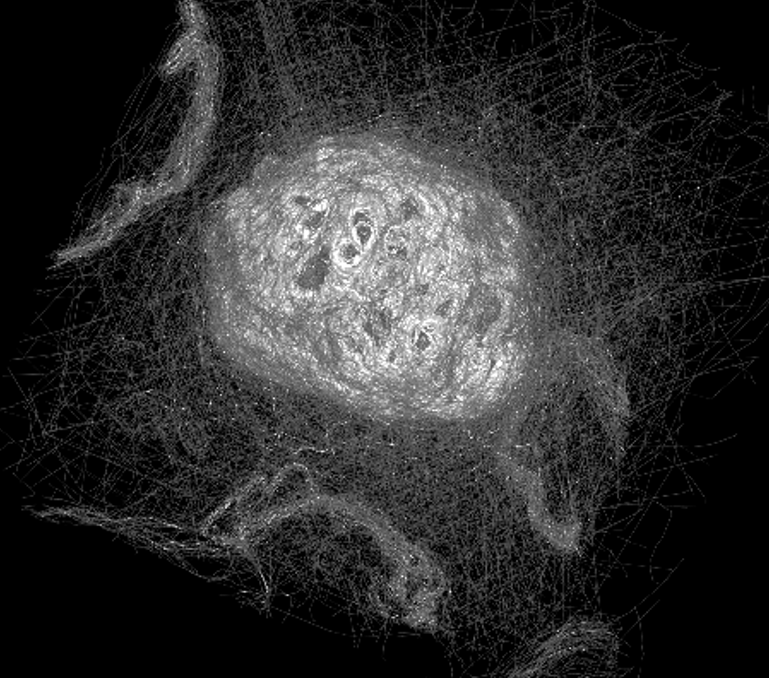
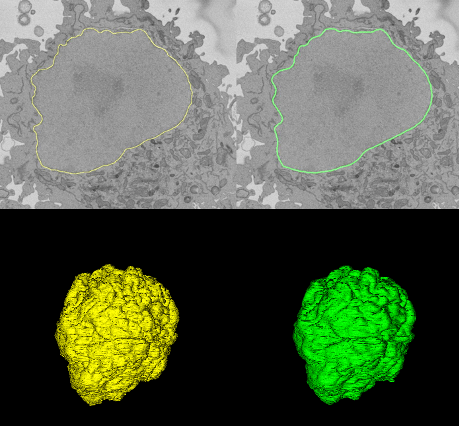
To make sense of all of this data, we developed an analysis approach that took all of these lines and averaged them to form a ‘consensus segmentation’ for each nuclear envelope. This consensus segmentation, produced through the collective effort of volunteers, was incredibly similar to that produced by an expert microscopist. You can see this in the image below: on the left (in yellow) you can see the expert segmentation of the nuclear envelope of one cell compared to the volunteer segmentation (in green). The top image shows a single slice from the cell, the bottom image shows the 3D reconstruction of the whole nuclear envelope.
4. Volunteer segmentations can be used to train powerful new algorithms capable of segmenting the nuclear envelope. We found that volunteer data alone, with no expert data at all, could be used to train computer algorithms to perform the task of nuclear envelope segmentation to a very high standard. In the gif below you can see the computer predicted nuclear envelope segmentation for each of the cells in pink.

5. Our algorithm works surprisingly well on other data sets. We ran this new algorithm on other datasets that had been produced under slightly different experimental conditions. Because of these differences, we didn’t expect the algorithm to perform very well, however, as you can see in the images below, it did a very good job at identifying the location of the nuclear envelope. Because of this transferability, members of our research team have already begun using this algorithm to aid their new research projects.

The future…
We’re so excited to share these results with you, our volunteer community, and the research communities we collaborate with, and we’re looking forward to building on these findings in the future. The algorithms we’ve been able to produce from this effort are already being used by research teams at the Crick, and we’ve already launched multiple new projects asking for your help to look at other organelles – The Etchiverse is expanding!
You can access all our current Etch A Cell projects through the Etch A Cell Organisation page
