Below is a guest post from Robbie Parks, a PhD student from Imperial College London who is studying how global climate change is influencing human mortality.
Read on to learn more about how crowd sourcing via platforms such as the Zooniverse can help us prepare for, and respond to, extreme weather.
– Helen
The future of extreme weather forecasting, preparation and response
Robbie Parks
The extreme weather around the globe during 2017 was a grim reminder of how truly devastating extreme weather can be. The citizens of Houston, New Orleans, Mumbai, Bangladesh, Nepal, Puerto Rico, the Dominican Republic, Florida and Sierra Leone joined the swelling ranks of victims from hurricanes and floods. Hurricane Harvey was followed by the powerful storms Irma, Jose, and Maria. Heatwaves and drought around the world, from India to Italy, have also brought misery and even death to millions.
When extreme weather strikes, lives and livelihoods are ruined, infrastructure destroyed, landscapes altered (sometimes irreversibly), and communities are often left with long-term mental and physical trauma. This clearly is a challenge not just for developing countries, but also for highly industrialised nations like the United States. What can we do about this?
Beyond avoiding the most devastating future extreme weather by curbing greenhouse gases improving forecast capabilities and capacity provides some of the best defence. National and local decision makers require reliable warnings to be able to provide emergency planning and relief in their locally minded extreme weather action plans.
Nowadays, the most modern supercomputers, armed with a suite of state-of-the art models of atmospheric processes, generate higher and higher spatial and temporal resolutions with steadily improving forecasting skill. Progress in this ‘quiet revolution’ of forecasting is significant.1 But the next 10 years could yet see another huge paradigm shift in forecast capabilities.
A critical part of this drive for improved weather forecasting is the World Weather Research Programme’s (WWRP) 10-year HIWeather (High Impact Weather) project, a consortium of over 2000 scientists from diverse fields participating from institutions worldwide. I spent some time with Dr Paolo Ruti, global chief of the WWRP at the World Meteorological Organization (WMO) in Geneva, Switzerland, to discuss advances in forecasting extreme weather through HIWeather.
According to Dr Ruti, the ambition of HIWeather is “to promote cooperative international research to achieve a dramatic increase in resilience to high impact weather”. This will translate to saving more lives on the ground in the onset of an extreme weather event, both by improved forecasting capability and better communication to decision makers who activate plans to manage such potential disasters.
First, Dr Ruti took me through the technological and analytical aspects of improving extreme weather forecasting. This includes advances in the resolution of the forecast models themselves, which enable the latest weather models to include physical processes which are critical to predicting extreme weather.
A good example (among many) of improved resolution making a difference to forecasting capability is models which allow for resolving convection processes (drawing moisture for hurricane generation at scales less than 10km spatially) to take place. The newest models are showing remarkable realism in their simulation of the potential evolution of severe storm events for up to 15 days. This additional forecast lead time and accuracy will become invaluable when planning evacuation in cities during hurricane season.
Another key component of improving forecasts is identifying what in weather patterns may indicate the onset of extremes. This includes the work of Dr Hannah Nissan, a researcher at the International Research Institute (IRI) for Climate and Society at Columbia University, New York. Dr Nissan mainly works with developing countries in Africa and South Asia to develop and improve extreme weather early warning systems. She is an expert at extending the forecast horizon of warnings by finding novel sources of predictability.
Dr Nissan and the IRI have researched the predictability of extreme heat in Bangladesh.2 Nissan has explored patterns of air transport and soil moisture, and found “heat waves are preceded by a characteristic wind pattern in the atmosphere, which can set itself up about a week to 10 days before a major heat wave.” Not only that, but reliable warnings up to 30 days in advance may be possible because as ‘soils are drier than normal for at least a month before a heat wave on average.” This will be essential to plan mid-term action plans in response to destructive heat waves.
Beyond improving modelling of weather and its extremes, challenges include data collection to power numerical and physical modelling of weather extremes. Data from the ground weather stations is the fuel for forecasting extreme weather. Developing countries like Bangladesh, however, suffer from a lack of essential weather measurement infrastructure. Dr Ruti of WWRP explains that “data, and real-time observations are a key challenge” for this purpose.
To address this challenge, Ruti said “weather data is being crowd sourced through third party networks using apps or portable weather measurement devices.” An example Ruti highlighted was of the use of mobile phones as a proxy for radar measurements to monitor and forecast rainfall. A pilot project by the French Institute for Development Research in Toulouse has demonstrated a proof-of-concept system in Burkina Faso, Niger, and Cameroon.3,4 This type of technology could emerge as especially important in parts of the world with poor ground-based weather monitoring infrastructure. With high penetration of mobile phones in most of the world,5 it is a promising example of where the next generation of weather data collection and assimilation into forecast models lies.
However, using data from public sources isn’t as simple as plugging it in to a forecasting system. Ruti warned me that “understanding the error characteristics of these data will be critical to using them effectively.” Prior to data post-processing using mobile phone data, the readings of rainfall could be two to three times the real value. Data needs to be trustworthy and processed correctly before it is used for forecast outcomes. Yet, the work is a positive move away from the expensive centralised government weather measurement projects in places which cannot necessarily afford to maintain them.
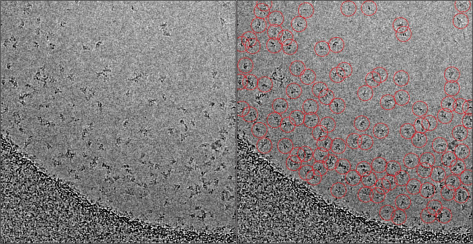
Crowd sourcing contributions in the aftermath of extreme weather is also a growing area. A leader in the field is Zooniverse, the world’s “largest and most popular platform for people-powered research.”6 The crowd sourcing of over 10,000 contributions to observing the devastation of Irma and Maria from satellite data has resulted in a damage analysis of buildings all over the Caribbean in just a few days, which would have traditionally taken a single researcher over a year. Partnering with the Machine Learning research group at Oxford, Zooniverse have built a powerful mechanism to catalogue damage from extreme weather events, which is invaluable for the clean-up and rebuilding phase.
Zooniverse continues to be involved with identifying the damage caused by the devastating hurricanes of summer 2017, as well as the historical classification of cyclones to help further the understanding of patterns and potential trends of cyclones under climate change. While these are two independent projects, they both serve the broad goal of increasing understanding and preparedness for any similar future events.
This kind of work attracts a wide range of contributors (based on a 2015 survey), with a balanced distribution of ages. Currently, those actively participating are mainly from English-speaking countries, as a combined 64% originate from the UK or the USA, with only 2% from developing countries. Many state that contributing to the projects on Zooniverse is fun, and that contributing to scientific progress is a strong part of the appeal.
Professor Brooke Simmons, Einstein fellow at UC San Diego, and leader of the Zooniverse Analysis Group, explains that the work users are doing on Zooniverse is a making a noticeable difference on the ground, with “really good feedback” from first responders, especially in response to Irma and Maria in 2017, where Zooniverse was called in to inspect the damage.
One such first responder, Rebekah Yore, of Rescue Global, an international Non-Governmental Organisation working to reduce disaster risk reduction around the world. Having previously worked on the response to Irma and Maria in 2017, Yore explains that Rescue Global could quickly find a set of personnel via Zooniverse whereby thousands of volunteers rapidly analysed pre- and post-storm satellite images to identify “specifically features of damage and hazard from infrastructure collapse, flooding and other causes”.
Once the volunteers’ work was done, the Machine Learning department at the University of Oxford would ‘run the responses through algorithms to filter anomalous results, improve the reliability of data, and produce heat maps of identified damage and features on the ground’. This became an essential component in Rescue Global’s humanitarian response, which was involved in determining which affected populations’ needs were the most critical, and which areas may have been more dangerous to traverse for teams deployed on the ground.
From a humanitarian response perspective, and particularly from the hurricane response in the Caribbean in 2017, Zooniverse allowed a great amount of accurate information to be generated extraordinarily quickly. Yore is quite clear that without Zooniverse, the first responders could not have had this kind of information to hand. Why was this information so useful? Yore sets the scene for what greets the those who are first to such disaster zones: “villages may have disappeared, there may be new, spontaneous groups of internally-displaced people. Bridges and access roads can also be destroyed, with flooding and landslides potentially blocking access”. The insights from Zooniverse are essential for both finding vulnerable populations while also protecting members of the rescue community.
Thus, arguably the most dramatic change to preparing for extreme weather is also how responses to them are managed. HIWeather has also identified endemic challenges of what a forecast should indicate, such as how to best spread effective warnings to a vulnerable community. Social media activity is a good way to analyse how effectively an extreme weather warning is being received and followed.7
While it is one thing to create experimental forecasting products; it is another to create something which decision-makers can use without requiring very specialised knowledge, often sorely lacking at Earth’s most vulnerable areas. Dr Joy Shumake-Guillemot leads the Joint Office for Climate and Health within the WMO, which specialises in translating forecasts into actions which local decision-makers can employ in extreme weather events.
Shumake-Guillemot recognizes how important it is to translate forecasting research into something useable for those on the ground: “the last-mile user (i.e. those who have to use the results for the purposes of minimising risk for vulnerable communities) in the ministry of health is often the neglected piece of the puzzle. Without building forecasting systems that factor in decision making, cutting-edge forecast products can so often become unused, especially in developing countries.”
Dr Ruti of WWRP also made clear to me that the key to saving lives and livelihoods is not only the forecasting technology, but also the methods of communicating the hazards to a community: ‘Better prediction and communication should go hand-in-hand’. We need to understand the ‘physical and social factors limiting the capability to communicate’, as well as understanding how to find better ways of forecasting.
A critical failure in the most devastating disaster to ever hit Myanmar, Cyclone Nargis in 2008, was that although the Indian Meteorological Office had predicted the extreme with four days’ notice,8 traditional methods of disseminating the warning (such as via TV and radio) did not reach isolated communities in the low-lying regions of the country, resulting in over 100,000 potentially avoidable deaths.
The link between decision makers and predictions often is at a disconnect, and this is the mid-term key to improving action on disaster forecasts. While events like Hurricane Harvey and others are truly terrible, they provide an opportunity to remind policy makers that it is critical we get the forecasting and communication coupling right. The future of forecasting includes targeted messaging to vulnerable populations, including the elderly, the young, prisoners, and labourers. Great gains can be made with existing technology and improved communication.
Of course, key steps forward in forecasting technology will occur over the next decade. But progress will not only develop in the models’ themselves, but also in the data collection for the models by crowd sourcing, and clarifying and communicating the forecasts to decision makers and then to vulnerable communities.
The future is full of extreme weather, but we can at least know a good deal more about it before it arrives.
References
1 Bauer P, Thorpe A, Brunet G. The quiet revolution of numerical weather prediction. Nature 2015; 525: 47–55.
2 Nissan H, Burkart K, Mason SJ, Coughlan de Perez E, van Aalst M. Defining and predicting heat waves in Bangladesh. J Appl Meteorol Climatol DOI:10.1175/JAMC-D-17-0035.1.
3 Tollefson J. Rain forecasts go mobile. Nature 2017; 544: 4–5.
4 Doumounia A, Gosset M, Cazenave F, Kacou M, Zougmore F. Rainfall monitoring based on microwave links from cellular telecommunication networks: First results from a West African test bed. Geophys Res Lett 2014; 41: 6015–21.
5 The World Bank. Mobile cellular subscriptions (per 100 people). 2017. https://data.worldbank.org/indicator/IT.CEL.SETS.P2?end=2016&start=1960&type=shaded&view=map&year_high_desc=true.
6 What is the Zooniverse? Zooniverse. 2017. https://www.zooniverse.org/about (accessed Sept 28, 2017).
7 Ripberger JT, Jenkins-Smith HC, Silva CL, Carlson DE, Henderson M. Social Media and Severe Weather: Do Tweets Provide a Valid Indicator of Public Attention to Severe Weather Risk Communication? Weather Clim Soc 2014; 6: 520–30.
8 IFRC. Community early warning systems. Guiding principles. 2012; : 84.